Example 2: Modelling with PyTracerLab in the Bayesian Statistical Framework¶
Here we show how to use the PyTracerLab package to perform a basic standard application of lumped parameter models (see Example 1), but with an important change. We still calibrate parameter values of the lumped parameter model from observations of tracer concentrations in an observation well, but here we perform this process in a Bayesian statistical framework using a Markov-chain Monte Carlo sampler.
import PyTracerLab.model as ism
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime
1. Load (Synthetic) Observation Data¶
See Example 5 on how this data is generated.
# load input series
# this would be the tracer concentration in precipitation or recharge in a
# practical problem
file_name = "example_input_series_1tracer.csv"
data = np.genfromtxt(
file_name,
delimiter=",",
dtype=["<U7", float],
encoding="utf-8",
skip_header=1
)
timestamps = np.array([datetime.strptime(row[0], r"%Y-%m") for row in data])
input_series = np.array([float(row[1]) for row in data])
# load observation series
# this would be the measured tracer concentration in groudnwater in a
# practical problem
file_name = "example_observation_series_1tracer.csv"
data = np.genfromtxt(
file_name,
delimiter=",",
dtype=["<U7", float],
encoding="utf-8",
skip_header=1
)
timestamps = np.array([datetime.strptime(row[0], r"%Y-%m") for row in data])
obs_series = np.array([float(row[1]) for row in data])
# load full system output series
# this would be the true tracer concentration in groudnwater in a practical
# problem; this is not available in practice
file_name = "example_output_series_1tracer.csv"
data = np.genfromtxt(
file_name,
delimiter=",",
dtype=["<U7", float],
encoding="utf-8",
skip_header=1
)
timestamps = np.array([datetime.strptime(row[0], r"%Y-%m") for row in data])
output_series = np.array([float(row[1]) for row in data])
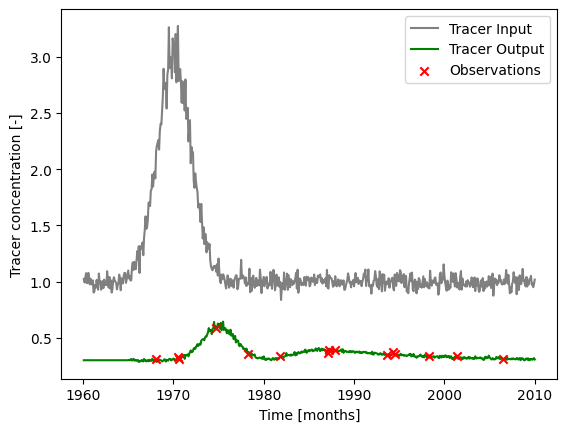
### plot input series, output series, and observations
# get observation timesteps
timesteps = [t.year + t.month / 12.0 for t in timestamps]
# create figure
fig, ax = plt.subplots(1, 1)
# plot input series
ax.plot(
timesteps,
input_series,
label="Tracer Input",
c="grey"
)
# plot output series
ax.plot(
timesteps,
output_series,
label="Tracer Output",
c="green"
)
# plot observations
ax.scatter(
timesteps,
obs_series,
label="Observations",
color="red",
marker="x",
zorder=10
)
ax.set_xlabel("Time [months]")
ax.set_ylabel("Tracer concentration [-]")
ax.legend()
plt.show()
2. Run MCMC for Model Parameters¶
t_half = 12.3 * 12.0
lambda_ = np.log(2.0) / t_half
### define model (use the same structure / units as the true model)
# time step is 1 month
m = ism.Model(
dt=1.0,
lambda_=lambda_,
input_series=input_series,
target_series=obs_series,
steady_state_input=1., # this is the true value
n_warmup_half_lives=10
)
# add an exponential-piston-flow unit
# define the initial model parameters for inference
epm_mtt_init = 12 * 40 # 1 year
epm_eta_init = 1.5
m.add_unit(
ism.EPMUnit(mtt=epm_mtt_init, eta=epm_eta_init),
fraction=.8, # 80 percent of the overall response; is the true value
bounds=[(12. * 39., 12.0 * 41.), (1.4, 1.6)],
prefix="epm"
)
# add a piston-flow unit
# define the true model parameters
pm_mtt_init = 12 * 5 # 1 year
m.add_unit(
ism.PMUnit(mtt=pm_mtt_init),
fraction=.2, # 20 percent of the overall response, is the true value
bounds=[(12. * 4, 12.0 * 6.)],
prefix="pm"
)
# create a solver
solver = ism.Solver(m)
# run MCMC
res = solver.dream_sample(
n_samples=2000, # effective samples after burn-in and thinning
burn_in=2000,
thin=1,
n_chains=6,
n_diff_pairs=2,
cr=[i / 5 for i in range(1, 6)],
sigma=0.01,
return_sim=True,
random_state=1234,
set_model_state=False
)
# print order of parameters in results dictionary
print(res["param_names"])
['epm.eta', 'epm.mtt', 'pm.mtt']
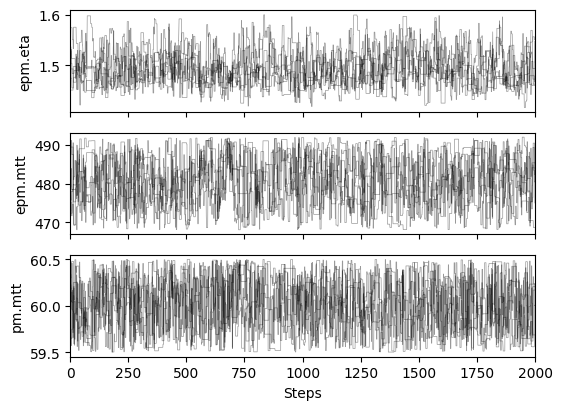
# plot chains to inspect convergence
fig, ax = plt.subplots(res["samples_chain"].shape[2], 1, figsize=(6, res["samples_chain"].shape[2] * 1.5), sharex=True)
for i in range(res["samples_chain"].shape[2]):
for j in range(res["samples_chain"].shape[0]):
ax[i].plot(res["samples_chain"][j, :, i], c="k", lw=.5, alpha=0.4)
ax[i].set_ylabel(m.param_keys(free_only=True)[i])
ax[i].set_xlim(0, res["samples_chain"].shape[1])
ax[-1].set_xlabel("Steps")
# print gelman rubin convergence statistics
print(res["gelman_rubin"])
{'epm.eta': 1.0076529086527881, 'epm.mtt': 1.0087310454766107, 'pm.mtt': 1.0036605287697642}
# define true parameters
epm_mtt_true = 12 * 40 # 40 years
epm_eta_true = 1.5
pm_mtt_true = 12 * 5 # 5 years (faster than EPM)
fig, ax = plt.subplots(1, 3, figsize=(10, 3))
_ = ax[0].hist(
res["samples"][:, 0],
bins=20,
density=True,
label="EPM Eta"
)
ax[0].axvline(epm_eta_true, color="red", label="True")
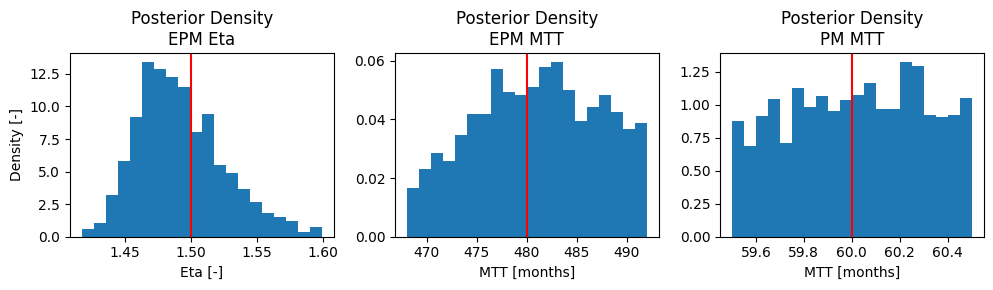
ax[0].set_title("Posterior Density\nEPM Eta")
ax[0].set_xlabel("Eta [-]")
ax[0].set_ylabel("Density [-]")
_ = ax[1].hist(
res["samples"][:, 1],
bins=20,
density=True,
label="EPM MTT"
)
ax[1].axvline(epm_mtt_true, color="red", label="True")
ax[1].set_title("Posterior Density\nEPM MTT")
ax[1].set_xlabel("MTT [months]")
_ = ax[2].hist(
res["samples"][:, 2],
bins=20,
density=True,
label="PM MTT"
)
ax[2].axvline(pm_mtt_true, color="red", label="True")
ax[2].set_title("Posterior Density\nPM MTT")
ax[2].set_xlabel("MTT [months]")
plt.tight_layout()
res["sims"].reshape(-1, res["sims"].shape[2]).shape
(12000, 600)
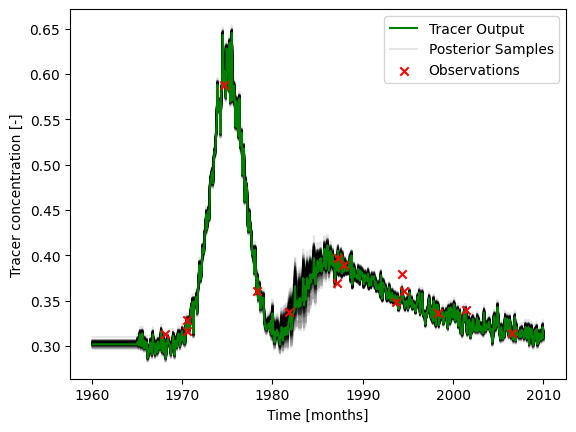
### plot input series, true output series, observations, and´
# calibrated output
# create figure
fig, ax = plt.subplots(1, 1)
# plot output series
ax.plot(
timesteps,
output_series,
label="Tracer Output",
c="green",
zorder=1000000
)
# plot calibrated results
sims_ = res["sims"].reshape(-1, res["sims"].shape[2])[::100, :]
for i in range(len(sims_)):
if i < len(sims_) - 1:
ax.plot(
timesteps,
sims_[i, :],
# label="Calibration",
c="black",
alpha=0.1
)
else:
ax.plot(
timesteps,
sims_[i, :],
label="Posterior Samples",
c="black",
alpha=0.1
)
# plot observations
ax.scatter(
timesteps,
obs_series,
label="Observations",
color="red",
marker="x",
zorder=100000000
)
ax.set_xlabel("Time [months]")
ax.set_ylabel("Tracer concentration [-]")
ax.legend()
plt.show()